AM
Pathogenes – Area 2
Antimicrobial Resistance (AM)
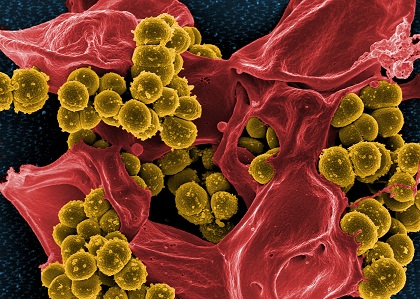
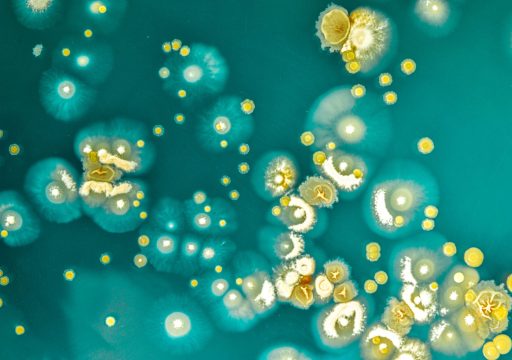
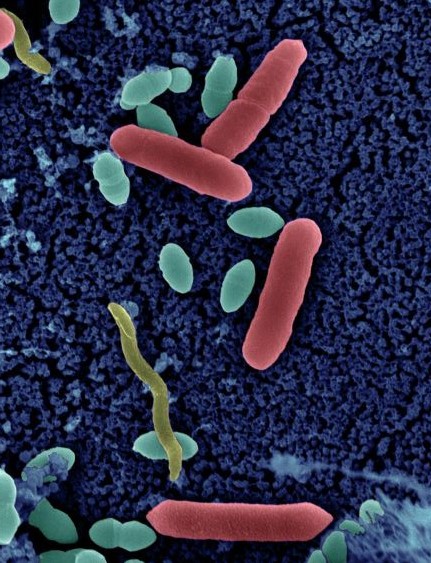
Escherichia coli, Bacteries
© Institut Pasteur
Analysis of clinical bacterial strains shows that the occurrence of AM can be associated with the overproduction or alteration of certain proteins as a result of mutations.
Identification of these stresses and the molecular responses triggered in PAs are essential for a better understanding of the environmental context that favours the occurrence and development of AM.
In addition to these intrinsic and adaptive (induced) resistance mechanisms, researchers in this topic are also interested in mechanisms acquired through the transfer of genetic information between pathogens, the latter posing a major health threat given their potential to spread.
As for fungal infections, the pressure exerted by the massive use of fungicides in agriculture is one of the main causes of cross-resistance with antifungals for medical use.
Given the current shortage of new antibiotics and antifungals, the detection and characterization of emerging resistance mechanisms, particularly to recently marketed molecules, is critical to rapidly identify the factors driving the emergence of resistance and provide information for the development of future molecules.