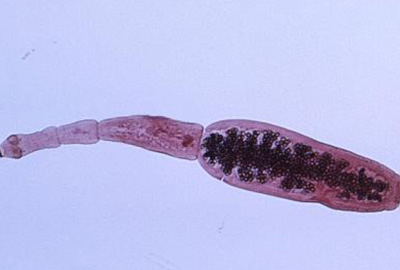
PATHOGÈNES
Thème Agents Pathogènes
Histoire, diffusion, résistance et génomique
Comment améliorer les traitements de demain ?
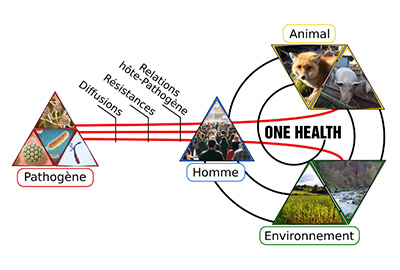
Les objectifs communs de ce thème sont de comprendre et d’identifier les facteurs favorisant l’émergence, la sélection et la diffusion des agents pathogènes. Les modifications de l’environnement, les interactions entre l’homme et l’animal, les contaminations par les agents sélecteurs de mutations et les transferts génétiques modifient ainsi l’histoire évolutive de ces pathogènes, à l’échelle de la population humaine, de la communauté microbienne (microbiotes) et de la cellule d’agent pathogène (single cell). La particularité de ce thème est d’avoir une approche intégrée, allant des observations (cliniques ou sur le terrain) et des caractérisations phénotypiques des agents pathogènes, jusqu’au décryptage des mécanismes moléculaires physiopathologiques à l’aide d’outils ‘omics’.
Coordinateurs du thème PATHOGENES :
- Frédéric Grenouillet, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENESfrederic.grenouillet@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau CHU Besançon
- Benoit Valot, Ingénieur de Recherche, PATHOGENES, PEA²t, Service qualitébenoit.valot@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau R217 (HdC)
Nos axes de recherche
Ces axes organisent la recherche “du terrain et de la clinique à la biologie moléculaire” sur des espèces microbiennes appartenant à tous les grands groupes d’AP (bactéries, parasites, levures, champignons, acariens et virus).
Diffusion des agents pathogènes humains
La diffusion des agents pathogènes (AP) est étudiée à partir de collections de micro-organismes isolés de prélèvements d’origine humaine, mais également environnementale (professionnel, domestique, hospitalier) et animale (élevage et faune sauvage).
Résistance aux agents antimicrobiens (RAM)
La lutte contre la résistance aux agents antimicrobiens est un défi majeur de santé publique et s’appuie sur l’étude de la prévalence et sur l’identification des déterminants géniques de résistance aux agents antibactériens et antifongiques.
Interaction hôte-pathogène
L’étude de la relation hôte-pathogène s’appuie sur une approche immunologique qui permet de quantifier l’exposition aux AP dans divers environnements par la mesure des anticorps produits (IgE pour les réponses allergiques immédiates et IgG pour les réponses infectieuses et immuno-allergiques).
Nos projets en cours
FLAMME
Le microbiote buccal est composé de 700 espèces de bactéries qui transforment les substances aromatiques. La composition de microbiote étant spécifique à son hôte, les mécanismes derrière le métabolisme microbien des substances aromatiques restent peu connus. FLAMME est un projet qui vise à comprendre le rôle du microbiote buccal dans la perception du goût des aliments en comparant le microbiote de participants ayant différents profils de perception du goût.
GAP-AFR (JPI-AMR)
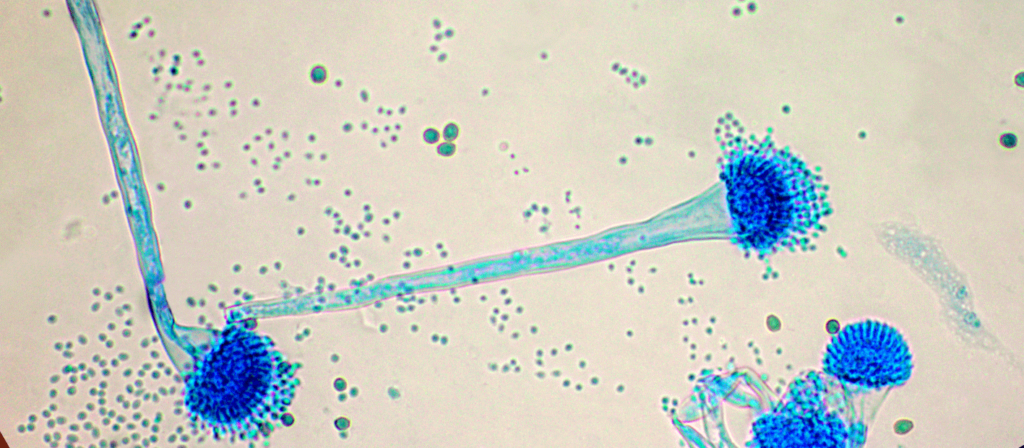
Les médicaments antifongiques azolés utilisés pour traiter les infections à Aspergillus fumigatus (pathogène fongique défini comme prioritaire par l’OMS) sont de moins en moins efficaces du fait de l’utilisation de molécules similaires pour la protection des cultures et des matériaux. Le projet vise à cartographier à l'échelle mondiale le pourcentage d’Aspergillus fumigatus résistants aux azolés, dont la croissance peut persister malgré le traitement antifongique.
MEHTA
Avec le changement climatique, une gestion raisonnée des ressources en eau devient cruciale pour l’irrigation. L’usage des eaux usées traitées s’impose en agriculture, mais les effets de l'ingestion de légumes irrigués par ces eaux sur le microbiote intestinal et la résistance aux antibiotiques restent inconnus. Pour y remédier, l’ANR/INSERM finance le projet MEHTA dans le cadre de France 2030, un programme de recherche prioritaire sur la résistance aux antibiotiques.
PARADE
Les phages sont des virus qui tuent spécifiquement les bactéries. Ils jouent un rôle crucial dans le contrôle des populations bactériennes dans les écosystèmes. Par un phénomène de coévolution, les bactéries ont développé une grande diversité de défenses anti-phages. Nous émettons l'hypothèse que le succès de certains clones épidémiques qui sont répandus à l'échelle mondiale doivent leur succès, au moins en partie, à leur plus grande résistance aux phages.
Nos actualités
Portrait de femme de science – Steffi Rocchi, microbiologiste
Un parcours inspirant mis en avant à l'occasion de la Journée internationale des filles et des femmes de science
Pourquoi huiles essentielles et antibiotiques ne font pas toujours bon ménage
Inhalées, ingérées, appliquées sur la peau… Les huiles essentielles agrémentent le quotidien de nombreuses personnes. Pourtant, ces substances ne sont pas anodines. Des travaux récents démontrent notamment que certaines d’entre elles peuvent interférer avec l’usage des antibiotiques.
Antifongiques, antibiotiques : des mécanismes de résistance identiques
Les mycoses résistantes aux médicaments antifongiques se multiplient. Notamment parce que les traitements contre les champignons pathogènes sont utilisés aussi bien en agriculture qu’en santé humaine et animale. Aspergillus fumigatus peut par exemple provoquer des...
Qui sommes-nous?
Coordinateurs⋅trices
- Frédéric Grenouillet, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENESfrederic.grenouillet@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau CHU Besançon
- Benoit Valot, Ingénieur de Recherche, PATHOGENES, PEA²t, Service qualitébenoit.valot@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau R217 (HdC)
Membres
- Victoria Aparicio, Doctorante, PATHOGENES, POLLUTIONvictoria.aparicio@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau 310K (La Bouloie)
- Coralie Barrera, Ingénieure de Recherche, PATHOGENES, PEA²tcoralie.barrera@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 34, bureau R28 (HdC)
- Rana Bazzi, Maîtresse de Conférences – UMLP, PATHOGENES, POLLUTIONrana.bazzi@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 66 62 94, bureau 308K (La Bouloie)
- Anne-Pauline Bellanger, Maître de conférences des universités - praticien hospitalier (MCU-PH) – UMLP, CHU Besançon, PATHOGENESanne-pauline.bellanger@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau CHU Besançon
- Xavier Bertrand, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENESxbertrand@-Code to remove to avoid SPAM-chu-besancon.fr, xbertran@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 70 63 21 36, bureau R+3 A8.1 (CHU)
- Adrien Biguenet, Membre associé – CHU Besançon, PATHOGENESadrien.biguenet@-Code to remove to avoid SPAM-univ-fcomte.fr, adrien.biguenet@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 70 63 21 46, bureau R216 (HdC)
- Louis Bohard, Doctorant, PATHOGENESlouis.bohard@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 21 91 83, bureau CHU Besançon
- Olympe Bonnardel, Doctorante, PATHOGENESolympe.bonnardel@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 75, bureau R245 (HdC)
- Kévin Bouiller, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENESkevin.bouiller@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 21 91 87, bureau CHU Besançon
- Thibault Bourdin, Postdoctorant, PATHOGENEStbourdin@-Code to remove to avoid SPAM-chu-besancon.fr, +33 (0)3 63 08 22 41, bureau R227 (HdC)
- Céline Bouvier-Slekovec, Maître de conférences des universités - praticien hospitalier (MCU-PH) – UMLP, CHU Besançon, PATHOGENESceline.bouvier@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 66 25 23 (fac) / +33 (0)3 70 63 25 97 (CHU), bureau S340 (UFR Santé), et bureau Service d’hygiène hospitalière (CHU Besançon)
- Bruno Cardey, Maître de Conférences – UMLP, PATHOGENES, POLLUTIONbruno.cardey@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 66 65 05, bureau 304K (La Bouloie)
- Audrey Cardot Laboissiere, Technicienne – CNRS, PATHOGENES, PEA²t, Service qualité, SERVICESaudrey.laboissiere@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau R217 (HdC), et bureau -211M (La Bouloie)
- Laura Camila Carrera Paéz, Postdoctorante, PATHOGENESlcarrerapaez@-Code to remove to avoid SPAM-chu-besancon.fr, +33 (0)3 63 08 22 41, bureau R227 (HdC)
- Catherine Chirouze, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENEScatherine.chirouze@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau CHU Besançon
- Pascal Cholley, Membre associé – Ingénieur d’étude et de recherche clinique CHU Besançon, PATHOGENESpcholley@-Code to remove to avoid SPAM-chu-besancon.fr
- Florent Demonmerot, Membre associé – Ingénieur d’étude et de recherche clinique CHU Besançon, PATHOGENESfdemonmerot@-Code to remove to avoid SPAM-chu-besancon.fr
- Nolwen Di Domizio, Doctorante, PATHOGENESnolwen.di_domizio@-Code to remove to avoid SPAM-edu.univ-fcomte.fr, +33 (0)3 63 08 22 15 , bureau R134 (HdC)
- Nathalie Floret, Maîtresse de conférences des universités - praticien hospitalier (MCU-PH) – UMLP, CHU Besançon, PATHOGENESNFloret@-Code to remove to avoid SPAM-chu-besancon.fr
- Adeline Gagnon, Doctorante, PATHOGENESadeline.gagnon@-Code to remove to avoid SPAM-umlp.fr, +33 (0)3 63 08 22 75, bureau R245 (HdC)
- Susie Gaillot, Ingénieure de Recherche – CHU Besançon, PATHOGENES+33 (0)3 70 63 21 68, bureau R214 (HdC)
- Patrick Giraudoux, Professeur Émérite, DYNABIO, PATHOGENESpatrick.giraudoux@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau -110M (La Bouloie)
- Mathieu Guillaume, Technicien – CHU Besançon, PATHOGENESm8guillaume@-Code to remove to avoid SPAM-chu-besancon.fr, +33 (0)3 63 08 22 33, bureau R37 (HdC)
- Audrey Guitton, Technicienne, PATHOGENES, PEA²t, Service qualitéaudrey.guitton@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 36, bureau R29 (HdC)
- Laura Henckel, Postdoctorante, DYNABIO, PATHOGENES, POLLUTION, SOPASTlaura.henckel@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 66 65 63, bureau -114M (La Bouloie)
- Didier Hocquet, Professeur – UMLP, PATHOGENESdhocquet@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 70 63 21 34, bureau R212 (HdC)
- Katy Jeannot, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENESkaty.jeannot@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau R213 (HdC)
- Clémence Kavira, Doctorante, PATHOGENESclemence.kavira@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau -109M (La Bouloie)
- Jenny Knapp, Ingénieure de Recherche – CHU Besançon, PATHOGENESjknapp@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 70 63 21 06, bureau R+3.B1.2 (CHU) / R251 (HdC)
- Quentin Lepiller, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENESquentin.lepiller@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau CHU Besançon
- Catherine Llanes, Professeure – UMLP, PATHOGENES, Service communicationchrono-env.direction@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 76, bureau R222 (HdC)
- Libor Makovicka, Professeur Émérite, PATHOGENESlibor.makovicka@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 99 46 74, bureau Montbéliard
- Tania Marx, Maîtresse de conférences des universités - praticien hospitalier (MCU-PH) – UMLP, CHU Besançon, PATHOGENEStania.marx@-Code to remove to avoid SPAM-univ-fcomte.fr
- Mathilde Massard, Maîtresse de Conférences – UMLP, PATHOGENESmathilde.massard@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 41, bureau R227 (HdC)
- Frédéric Mauny, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENESfrederic.mauny@-Code to remove to avoid SPAM-univ-fcomte.fr
- Laurence Millon, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENESlmillon@-Code to remove to avoid SPAM-chu-besancon.fr, +33 (0)3 70 63 23 53, bureau CHU Besançon
- Eloïse Paulet, Doctorante, PATHOGENESeloise.paulet@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 75, bureau R245 (HdC)
- Antoine Perasso, Professeur – UMLP, DYNABIO, GEODE, PATHOGENES, POLLUTIONantoine.perasso@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau -110L (La Bouloie)
- Anaïs Potron, Maître de conférences des universités - praticien hospitalier (MCU-PH) – UMLP, CHU Besançon, PATHOGENESanais.potron@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 70 63 21 68, bureau R214 (HdC)
- Jean-Luc Prétet, Professeur des universités - praticien hospitalier (PU-PH) – UMLP, CHU Besançon, PATHOGENESjean_luc.pretet@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 70 63 20 51, bureau HdC
- Emma Prétot, Doctorante, CHU Besançon, PATHOGENESemma.pretot@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau R245 (HdC)
- Francis Raoul, Professeur – UMLP, DYNABIO, PATHOGENES, POLLUTIONfrancis.raoul@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 66 57 36, bureau -114L (La Bouloie)
- Dominique Rieffel, Technicien, PATHOGENES, PEA²t, POLLUTIONdominique.rieffel@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 66 63 52, bureau -104L (La Bouloie)
- Steffi Rocchi, Chercheuse associée, PATHOGENES, POLLUTIONsteffi.rocchi@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 39, bureau R210 (HdC)
- Kévin Roche, Postdoctorant, PATHOGENES, PEA²t, SOPASTkevin.roche@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau -213M (La Bouloie)
- Julie Rousselot, Doctorante, CHU Besançon, PATHOGENESjulie.rousselot@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 33, bureau R27 (HdC)
- Stéphane Roux, Professeur – UMLP, PATHOGENES, POLLUTIONstephane.roux@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 66 62 99, bureau 308K (La Bouloie)
- Adeline Rouzet-San, Ingénieure de Recherche – CHU Besançon (Parasitologie-Mycologie), PATHOGENES, PEA²tadeline.rouzet@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 34, bureau R28 (HdC)
- Lison Schmidlin, Adjointe Technique de Recherche et de Formation (ITRF) – UFR Santé, PATHOGENES, PEA²tlison.schmidlin@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 63 08 22 36, bureau R29 (HdC)
- Hugo Sentenac, Maître de Conférences – IUT Besançon-Vesoul, DYNABIO, PATHOGENES, POLLUTIONhugo.sentenac@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau -105M (La Bouloie)
- Momar Thiaw Sylla, Doctorant, PATHOGENESmomar.thiaw_sylla@-Code to remove to avoid SPAM-edu.univ-fcomte.fr, bureau R212 (HdC)
- Freddy Torrealba Anzola, Ingénieur de Recherche – UMLP, PATHOGENES, PEA²tfreddy.torrealba_anzola@-Code to remove to avoid SPAM-univ-fcomte.fr, +33 (0)3 81 66 65 17, bureau 309K (La Bouloie)
- Victor Zosim, Doctorant, CHU Besançon, PATHOGENESvictor.zosim@-Code to remove to avoid SPAM-univ-fcomte.fr, bureau HdC

Laboratoire Chrono-environnement
UMR 6249
Université Marie et Louis Pasteur
16, Route de Gray
25030 Besançon cedex
Téléphone : +33 (0)3 81 66 62 55
Télécopie : +33 (0)3 81 66 65 68
Tous droits réservés chrono-environnement | Mentions légales | …